Abstract
A major gap in the Plasmodium vivax elimination toolkit is the identification of individuals carrying clinically silent and undetectable liver-stage parasites, called hypnozoites. This study developed a panel of serological exposure markers capable of classifying individuals with recent P. vivax infections who have a high likelihood of harboring hypnozoites. We measured IgG antibody responses to 342 P. vivax proteins in longitudinal clinical cohorts conducted in Thailand and Brazil and identified candidate serological markers of exposure. Candidate markers were validated using samples from year-long observational cohorts conducted in Thailand, Brazil and the Solomon Islands and antibody responses to eight P. vivax proteins classified P. vivax infections in the previous 9 months with 80% sensitivity and specificity. Mathematical models demonstrate that a serological testing and treatment strategy could reduce P. vivax prevalence by 59–69%. These eight antibody responses can serve as a biomarker, identifying individuals who should be targeted with anti-hypnozoite therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data and code availability
All data and code for reproducing this analysis are available online at https://github.com/MWhite-InstitutPasteur/Pvivax_sero_dx.
References
Sutanto, I. et al. Negligible impact of mass screening and treatment on mesoendemic malaria transmission at West Timor in eastern Indonesia: a cluster-randomized trial. Clin. Infect. Dis. 67, 1364–1372 (2018).
Waltmann, A. et al. High rates of asymptomatic, sub-microscopic Plasmodium vivax infection and disappearing Plasmodium falciparum malaria in an area of low transmission in Solomon Islands. PLoS Negl. Trop. Dis. 9, e0003758 (2015).
Mueller, I. et al. Key gaps in the knowledge of Plasmodium vivax, a neglected human malaria parasite. Lancet Infect. Dis. 9, 555–566 (2009).
Moreira, C. M., Abo-Shehada, M., Price, R. N. & Drakeley, C. J. A systematic review of sub-microscopic Plasmodium vivax infection. Malar. J. 14, 360 (2015).
Robinson, L. J. et al. Strategies for understanding and reducing the Plasmodium vivax and Plasmodium ovale hypnozoite reservoir in Papua New Guinean children: a randomised placebo-controlled trial and mathematical model. PLoS Med. 12, e1001891 (2015).
Parker, D. M. et al. Microgeography and molecular epidemiology of malaria at the Thailand-Myanmar border in the malaria pre-elimination phase. Malar. J. 14, 198 (2015).
Mogeni, P. et al. Detecting malaria hotspots: a comparison of rapid diagnostic test, microscopy, and polymerase chain reaction. J. Infect. Dis. 216, 1091–1098 (2017).
Ding, X. C. et al. Defining the next generation of Plasmodium vivax diagnostic tests for control and elimination: target product profiles. PLoS Negl. Trop. Dis. 11, e0005516 (2017).
World Health Organization. Malaria Surveillance, Monitoring & Evaluation: A Reference Manual (2018).
Harris, I. et al. A large proportion of asymptomatic Plasmodium infections with low and sub-microscopic parasite densities in the low transmission setting of Temotu Province, Solomon Islands: challenges for malaria diagnostics in an elimination setting. Malar. J. 9, 254 (2010).
Sutanto, I. et al. Negligible impact of mass screening and treatment on meso-endemic malaria transmission at West Timor in eastern Indonesia: a cluster-randomised trial. Clin. Infect. Dis. 67, 1364–1372 (2018).
Ong, K. I. C. et al. Systematic review of the clinical manifestations of glucose-6-phosphate dehydrogenase deficiency in the Greater Mekong subregion: implications for malaria elimination and beyond. BMJ Glob. Health 2, e000415 (2017).
Longley, R. J. et al. Asymptomatic Plasmodium vivax infections induce robust IgG responses to multiple blood-stage proteins in a low-transmission region of western Thailand. Malar. J. 16, 178 (2017).
Longley, R. J. et al. Acquisition and longevity of antibodies to Plasmodium vivax preerythrocytic antigens in western Thailand. Clin. Vaccine Immunol. 23, 117–124 (2016).
Helb, D. A. et al. Novel serologic biomarkers provide accurate estimates of recent Plasmodium falciparum exposure for individuals and communities. Proc. Natl Acad. Sci. USA 112, E4438–E4447 (2015).
Welch, R. J., Anderson, B. L. & Litwin, C. M. Rapid immunochromatographic strip test for detection of anti-K39 immunoglobulin G antibodies for diagnosis of visceral leishmaniasis. Clin. Vaccine Immunol. 15, 1483–1484 (2008).
Gwyn, S. et al. Lateral flow-based antibody testing for Chlamydia trachomatis. J. Immunol. Methods 435, 27–31 (2016).
Landry, M. L. Diagnostic tests for influenza infection. Curr. Opin. Pediatr. 23, 91–97 (2011).
Greenhouse, B., Smith, D. L., Rodriguez-Barraquer, I., Mueller, I. & Drakeley, C. J. Taking sharper pictures of malaria with CAMERAs: combined antibodies to measure exposure recency assays. Am. J. Trop. Med. Hyg. 99, 1120–1127 (2018).
White, N. J. Determinants of relapse periodicity in Plasmodium vivax malaria. Malar. J. 10, 297 (2011).
Battle, K. E. et al. Geographical variation in Plasmodium vivax relapse. Malar. J. 13, 144 (2014).
White, M. T., Shirreff, G., Karl, S., Ghani, A. C. & Mueller, I. Variation in relapse frequency and the transmission potential of Plasmodium vivax malaria. Proc. Biol. Sci. 283, 20160048 (2016).
Longley, R. J. et al. Naturally acquired antibody responses to more than 300 Plasmodium vivax proteins in three geographic regions. PLoS Negl. Trop. Dis. 11, e0005888 (2017).
Tsuboi, T. et al. Wheat germ cell-free system-based production of malaria proteins for discovery of novel vaccine candidates. Infect. Immun. 76, 1702–1708 (2008).
Franca, C. T. et al. Identification of highly-protective combinations of Plasmodium vivax recombinant proteins for vaccine development. eLife 6, e28673 (2017).
Franca, C. T. et al. Plasmodium vivax reticulocyte binding proteins are key targets of naturally acquired immunity in young Papua New Guinean children. PLoS Negl. Trop. Dis. 10, e0005014 (2016).
Lu, F. et al. Profiling the humoral immune responses to Plasmodium vivax infection and identification of candidate immunogenic rhoptry-associated membrane antigen (RAMA). J. Proteomics 102, 66–82 (2014).
Franca, C. T. et al. An antibody screen of a Plasmodium vivax antigen library identifies novel merozoite proteins associated with clinical protection. PLoS Negl. Trop. Dis. 10, e0004639 (2016).
Gruszczyk, J. et al. Transferrin receptor 1 is a reticulocyte-specific receptor for Plasmodium vivax. Science 359, 48–55 (2018).
Soares, I. S. et al. A Plasmodium vivax vaccine candidate displays limited allele polymorphism, which does not restrict recognition by antibodies. Mol. Med. 5, 459–470 (1999).
Wang, B. et al. Immunoprofiling of the tryptophan-rich antigen family in Plasmodium vivax. Infect. Immun. 83, 3083–3095 (2015).
Cheng, Y. et al. Naturally acquired humoral and cellular immune responses to Plasmodium vivax merozoite surface protein 8 in patients with infection. Malar. J. 16, 211 (2017).
Hester, J. et al. De novo assembly of a field isolate genome reveals novel Plasmodium vivax erythrocyte invasion genes. PLoS Negl. Trop. Dis. 7, e2569 (2013).
Jiang, J., Barnwell, J. W., Meyer, E. V. & Galinski, M. R. Plasmodium vivax merozoite surface protein-3 (PvMSP3): expression of an 11 member multigene family in blood-stage parasites. PLoS ONE 8, e63888 (2013).
Ntege, E. H. et al. Blood-stage malaria vaccines: post-genome strategies for the identification of novel vaccine candidates. Expert Rev. Vaccines 16, 769–779 (2017).
White, M. T. et al. Mathematical modelling of the impact of expanding levels of malaria control interventions on Plasmodium vivax. Nat. Commun. 9, 3300 (2018).
Rabinovich, R. N. et al. malERA: an updated research agenda for malaria elimination and eradication. PLoS Med. 14, e1002456 (2017).
Cook, J. et al. Using serological measures to monitor changes in malaria transmission in Vanuatu. Malar. J. 9, 169 (2010).
Ladeia-Andrade, S. et al. Naturally acquired antibodies to merozoite surface protein (MSP)-1(19) and cumulative exposure to Plasmodium falciparum and Plasmodium vivax in remote populations of the Amazon Basin of Brazil. Mem. Inst. Oswaldo Cruz 102, 943–951 (2007).
Cook, J. et al. Sero-epidemiological evaluation of changes in Plasmodium falciparum and Plasmodium vivax transmission patterns over the rainy season in Cambodia. Malar. J. 11, 86 (2012).
White, M. T. et al. Plasmodium vivax and Plasmodium falciparum infection dynamics: re-infections, recrudescences and relapses. Malar. J. 17, 170 (2018).
White, M. T. et al. Dynamics of the antibody response to Plasmodium falciparum infection in African children. J. Infect. Dis. 210, 1115–1122 (2014).
Boyle, M. J., Reiling, L., Osier, F. H. & Fowkes, F. J. Recent insights into humoral immunity targeting Plasmodium falciparum and Plasmodium vivax malaria. Int. J. Parasitol. 47, 99–104 (2017).
Richards, J. S. et al. Association between naturally acquired antibodies to erythrocyte-binding antigens of Plasmodium falciparum and protection from malaria and high-density parasitemia. Clin. Infect. Dis. 51, e50–e60 (2010).
Arama, C. et al. Genetic resistance to malaria is associated with greater enhancement of immunoglobulin (Ig)M than IgG responses to a broad array of Plasmodium falciparum antigens. Open Forum Infect. Dis. 2, ofv118 (2015).
Woodberry, T. et al. Antibodies to Plasmodium falciparum and Plasmodium vivax merozoite surface protein 5 in Indonesia: species-specific and cross-reactive responses. J. Infect. Dis. 198, 134–142 (2008).
Douglas, N. M. et al. Plasmodium vivax recurrence following falciparum and mixed species malaria: risk factors and effect of antimalarial kinetics. Clin. Infect. Dis. 52, 612–620 (2011).
Looareesuwan, S., White, N. J., Chittamas, S., Bunnag, D. & Harinasuta, T. High rate of Plasmodium vivax relapse following treatment of falciparum malaria in Thailand. Lancet 2, 1052–1055 (1987).
Fornace, K. M. et al. Exposure and infection to Plasmodium knowlesi in case study communities in Northern Sabah, Malaysia and Palawan, the Philippines. PLoS Negl. Trop. Dis. 12, e0006432 (2018).
Herman, L. S. et al. Identification and validation of a novel panel of Plasmodium knowlesi biomarkers of serological exposure. PLoS Negl. Trop. Dis. 12, e0006457 (2018).
White, N. J. & Imwong, M. Relapse. Adv. Parasitol. 80, 113–150 (2012).
Alonso, P. L. et al. A research agenda to underpin malaria eradication. PLoS Med. 8, e1000406 (2011).
Hietanen, J. et al. Gene models, expression repertoire, and immune response of Plasmodium vivax reticulocyte binding proteins. Infect. Immun. 84, 677–685 (2015).
Cole-Tobian, J. L. et al. Strain-specific duffy binding protein antibodies correlate with protection against infection with homologous compared to heterologous Plasmodium vivax strains in Papua New Guinean children. Infect. Immun. 77, 4009–4017 (2009).
Nguitragool, W. et al. Highly heterogeneous residual malaria risk in western Thailand. Int. J. Parasitol. 49, 455–462 (2019).
Quah, Y. W. et al. Molecular epidemiology of residual Plasmodium vivax transmission in a paediatric cohort in Solomon Islands. Malar. J. 18, 106 (2019).
Wampfler, R. et al. Strategies for detection of Plasmodium species gametocytes. PLoS ONE 8, e76316 (2013).
Rosanas-Urgell, A. et al. Comparison of diagnostic methods for the detection and quantification of the four sympatric Plasmodium species in field samples from Papua New Guinea. Malar. J. 9, 361 (2010).
Healer, J. et al. Vaccination with conserved regions of erythrocyte-binding antigens induces neutralizing antibodies against multiple strains of Plasmodium falciparum. PLoS ONE 8, e72504 (2013).
Borremans, B., Hens, N., Beutels, P., Leirs, H. & Reijniers, J. Estimating time of infection using prior serological and individual information can greatly improve incidence estimation of human and wildlife infections. PLoS Comput. Biol. 12, e1004882 (2016).
Kirkpatrick, S., Gelatt, C. D. Jr. & Vecchi, M. P. Optimization by simulated annealing. Science 220, 671–680 (1983).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction (Springer, 2009).
Acknowledgments
We acknowledge the extensive field teams that contributed to sample collection and qPCR assays, including A. Kuehn, Y. W. Quah, P. Sripoorote and A. Waltmann. We thank all the individuals who participated in each of the studies and thank the Australian and Thai Red Cross and the RBB for donation of plasma samples. We thank the Volunteer Blood Donor Registry at WEHI for donation of plasma samples and L. Laskos and J. Harris for their collection and advice. We thank C. King (Case Western Reserve University) for provision of the Papua New Guinea control plasma pool. M. Bahlo (Walter and Eliza Hall Institute) is thanked for her advice on algorithm development. We acknowledge funding from the Global Health Innovative Technology Fund (T2015-142 to I.M.), the National Institute of Allergy and Infectious Diseases (National Institutes of Health grant 5R01 AI 104822 to J.S. and 5U19AI089686-06 to J.K.) and the National Health and Medical Research Council Australia (1092789 and 1134989 to I.M. and 1143187 to W.-H.T.). Cohort samples were derived from field studies originally funded by the TransEPI consortium (supported by the Bill and Melinda Gates Foundation). This work has been supported by FIND with funding from the Australian and British governments. We also acknowledge support from the National Research Council of Thailand. This work was made possible through Victorian State Government Operational Infrastructure Support and Australian Government National Health and Medical Research Council (NHMRC) Independent Research Institute Infrastructure Support Scheme. I.M. is supported by an NHMRC Senior Research Fellowship (1043345). D.L.D. is supported by an NHMRC Principal Research Fellowship (1023636). T.T. was supported in part by Japan Society for the Promotion of Science Grant-in-Aid for Scientific Research (KAKENHI, JP15H05276, JP16K15266). W.H.T. is a Howard Hughes Medical Institute–Wellcome Trust International Research Scholar (208693/Z/17/Z). R.J.L. received the Page Betheras Award from WEHI to provide funding for technical support for this project during parental leave. M.V.G.L. and W.M.M. are fellows of the Brazilian National Council for Scientific and Technological Development.
Author information
Authors and Affiliations
Contributions
R.J.L., M.T.W., T.T. and I.M. designed the study. W.N., W.M., J.K., M.L., J.S. and I.M. conducted the cohort studies. E.T., M.M., M.H., F.M., W.-H.T., J.H., C.H., C.E.C. and T.T. expressed the proteins. R.J.L., E.T., M.M., J.B. and Z.S.-J.L. performed antibody measurements. R.J.L., C.S.N.L.-W.-S. and M.T.W. conducted data management and analysis. M.T.W. and T.O. performed modeling. R.J.L., M.T.W. and I.M. wrote the draft of the report. L.J.R., C.P., D.L.D., X.C.D. and I.J.G. provided expert advice. All authors contributed to data interpretation and revision of the report.
Corresponding author
Ethics declarations
Competing interests
FIND contributed to early funding of this work and had a role in data interpretation, writing of the report and the decision to submit. No other funders of this study had any role in study design, data collection, data analysis, data interpretation, writing of the report or the decision to submit. R.J.L., M.W., T.T. and I.M. are inventors on patent PCT/US17/67926 on a system, method, apparatus and diagnostic test for P. vivax.
Additional information
Peer review information Alison Farrell is the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Study design and follow-up schedule.
a, Thai and Brazilian patients were enrolled following a clinical episode of P. vivax and treated according to the relevant National Guidelines, with directly observed treatment (DOT) to ensure compliance and reduced risk of relapse. Volunteers were followed for nine months after enrollment, with finger-prick blood samples collected at enrollment and week 1, then every two weeks for six months, then every month. Antibody levels were measured in a subset of 32 Thai and 33 Brazilian volunteers who were confirmed to be free of blood-stage Plasmodium parasites by analysing all samples by light microscopy and qPCR. b, 999 participants from Thailand, 1274 participants from Brazil, and 860 participants from the Solomon Islands were followed longitudinally for 12 months with active surveillance visits every month. For the analysis in the validation phase antibody levels were measured in plasma samples from the last visit. For the analysis in the application phase, antibody levels were measured in plasma samples from the first visit.
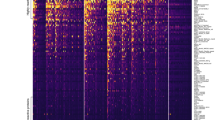
Extended Data Fig. 2 Measured antibody responses to 60 proteins on the Luminex® platform, stratified by geographical location and time since last PCR detected infection.
Antibody responses to 60 antigens measured in n = 2,281 biologically independent samples on the Luminex® platform, stratified by geographical location and time since last PCR detected infection. Boxplots denote median and interquartile ranges (IQR) of the data, with whiskers denoting the median ± 1.5*IQR.
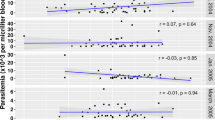
Extended Data Fig. 3 Association between background reactivity in non-malaria exposed controls and ranking of candidate SEMs by area under the curve (AUC).
Mean relative antibody units (RAU) detected in malaria-naïve control panels from Melbourne, Australia (n=202), Bangkok, Thailand (n=72) and Rio de Janeiro Brazil (n = 96) compared to the AUC of the 60 candidate P.vivax proteins generated during the validation phase. WGCF expressed proteins are in black and E. coli or Baculovirus expressed proteins are in blue. RBP2b161-1454 (E. coli) is in red and RBP2b1986-2653 is in orange.
Extended Data Fig. 4 Breakdown of the classification of the Random Forests algorithm with target sensitivity and specificity of 80%.
The size of each rectangle is proportional to the number of samples in each category (See Supplementary Table 4 of accompanying manuscript). The coloured area represents the proportion correctly classified, and the shaded area represents the proportion mis-classified.
Extended Data Fig. 5 Receiver operating characteristic (ROC) curve for the composite classification algorithm.
All curves presented are the median of 1000 repeat cross-validations.
Supplementary information
Supplementary Information
Supplementary Methods and Results, Supplementary Figs. 1–12 and Tables 1–4.
Rights and permissions
About this article
Cite this article
Longley, R.J., White, M.T., Takashima, E. et al. Development and validation of serological markers for detecting recent Plasmodium vivax infection. Nat Med 26, 741–749 (2020). https://doi.org/10.1038/s41591-020-0841-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-0841-4
This article is cited by
-
Comparing malaria risk exposure in rural Cambodia population using GPS tracking and questionnaires
Malaria Journal (2024)
-
Antibody selection strategies and their impact in predicting clinical malaria based on multi-sera data
BioData Mining (2024)
-
Using serological diagnostics to characterize remaining high-incidence pockets of malaria in forest-fringe Cambodia
Malaria Journal (2024)
-
Malaria seroepidemiology in very low transmission settings in the Peruvian Amazon
Scientific Reports (2024)
-
Serological evaluation of risk factors for exposure to malaria in a pre-elimination setting in Malaysian Borneo
Scientific Reports (2023)