Abstract
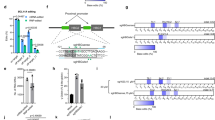
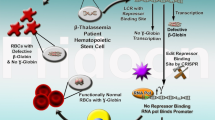
Gene editing to disrupt the GATA1-binding site at the +58 BCL11A erythroid enhancer could induce γ-globin expression, which is a promising therapeutic strategy to alleviate β-hemoglobinopathy caused by HBB gene mutation. In the present study, we report the preliminary results of an ongoing phase 1/2 trial (NCT04211480) evaluating safety and efficacy of gene editing therapy in children with blood transfusion-dependent β-thalassemia (TDT). We transplanted BCL11A enhancer-edited, autologous, hematopoietic stem and progenitor cells into two children, one carrying the β0/β0 genotype, classified as the most severe type of TDT. Primary endpoints included engraftment, overall survival and incidence of adverse events (AEs). Both patients were clinically well with multilineage engraftment, and all AEs to date were considered unrelated to gene editing and resolved after treatment. Secondary endpoints included achieving transfusion independence, editing rate in bone marrow cells and change in hemoglobin (Hb) concentration. Both patients achieved transfusion independence for >18 months after treatment, and their Hb increased from 8.2 and 10.8 g dl−1 at screening to 15.0 and 14.0 g dl−1 at the last visit, respectively, with 85.46% and 89.48% editing persistence in bone marrow cells. Exploratory analysis of single-cell transcriptome and indel patterns in edited peripheral blood mononuclear cells showed no notable side effects of the therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All amplicon deep-sequencing data generated in this article can be found at the National Center for Biotechnology Information’s Sequence Read Archive (accession no. PRJNA839164). Single-cell transcriptome data generated in this article can be found at the Gene Expression Omnibus database with accession no. GSE204688. A shared dataset used in this article for 10,000 PBMC single-cell transcriptome data from a healthy donor can be downloaded from: https://cf.10xgenomics.com/samples/cell-exp/4.0.0/SC3_v3_NextGem_SI_PBMC_10K/SC3_v3_NextGem_SI_PBMC_10K_raw_feature_bc_matrix.tar.gz.
Individual participant data (IPD) that underlie the results reported in published article will be shared, after de-identification (text, tables, figures and appendices). Other available documents include the study protocol. IPD sharing will start at 6 months and end at 36 months after article publication. IPD will be shared with investigators for individual data meta-analysis, after their proposed use of the data has been approved by an independent review committee. Proposals should be directed to yxwu@bio.ecnu.edu.cn and fu.bin@csu.edu.cn. To gain access, data requestors will need to sign a data access agreement. Source data are provided with this paper.
References
Piel, F. B. The present and future global burden of the inherited disorders of hemoglobin. Hematol. Oncol. Clin. 30, 327–341 (2016).
Cao, A. & Galanello, R. Beta-thalassemia. Genet. Med. 12, 61–76 (2010).
Taher, A. T., Weatherall, D. J. & Cappellini, M. D. Thalassaemia. Lancet 391, 155–167 (2018).
Thein, S. L. The molecular basis of β-thalassemia. Cold Spring Harb. Perspect. Med. 3, a011700 (2013).
Yin, X. L. et al. Treatment and complications of thalassemia major in Guangxi, Southern China. Pediatr. Blood Cancer 57, 1174–1178 (2011).
Modell, B., Khan, M. & Darlison, M. Survival in β-thalassaemia major in the UK: data from the UK Thalassaemia Register. Lancet 355, 2051–2052 (2000).
Li, C. et al. Related and unrelated donor transplantation for β-thalassemia major: results of an international survey. Blood Adv. 3, 2562–2570 (2019).
Sankaran, V. G. & Orkin, S. H. The switch from fetal to adult hemoglobin. Cold Spring Harb. Perspect. Med 3, a011643–a011643 (2013).
Bauer, D. E. et al. An erythroid enhancer of BCL11A subject to genetic variation determines fetal hemoglobin level. Science 342, 253–257 (2013).
Bauer, D. E. & Orkin, S. H. Hemoglobin switching’s surprise: the versatile transcription factor BCL11A is a master repressor of fetal hemoglobin. Curr. Opin. Genet Dev. 33, 62–70 (2015).
Uda, M. et al. Genome-wide association study shows BCL11A associated with persistent fetal hemoglobin and amelioration of the phenotype of beta-thalassemia. Proc. Natl Acad. Sci. USA 105, 1620–1625 (2008).
Demirci, S. et al. BCL11A enhancer-edited hematopoietic stem cells persist in rhesus monkeys without toxicity. J. Clin. Invest. 130, 6677–6687 (2020).
Brendel, C. et al. Lineage-specific BCL11A knockdown circumvents toxicities and reverses sickle phenotype. J. Clin. Invest. 126, 3868–3878 (2016).
Wu, Y. et al. Highly efficient therapeutic gene editing of human hematopoietic stem cells. Nat. Med. 25, 776–783 (2019).
Brendel, C. et al. Preclinical evaluation of a novel lentiviral vector driving lineage-specific BCL11A knockdown for sickle cell gene therapy. Mol. Ther. Methods Clin. Dev. 17, 589–600 (2020).
Esrick, E.B. et al. Post-transcriptional genetic silencing of BCL11A to treat sickle cell disease. N. Engl. J. Med. 384, 284–285 (2021).
Canver, M. C. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015).
Frangoul, H. et al. CRISPR–Cas9 gene editing for sickle cell disease and β-thalassemia. N. Engl. J. Med. 384, 252–260 (2021).
Wang, L. et al. Reactivation of γ-globin expression through Cas9 or base editor to treat β-hemoglobinopathies. Cell Res. 30, 276–278 (2020).
Gonçalves, T. L., Benvegnú, D. M. & Bonfanti, G. Specific factors influence the success of autologous and allogeneic hematopoietic stem cell transplantation. Oxid. Med. Cell Longev. 2, 82–87 (2009).
Jillella, A., Kallab, A. & Kutlar, A. Autoimmune thrombocytopenia following autologous hematopoietic cell transplantation: review of literature and treatment options. Bone Marrow Transplant. 26, 925–927 (2000).
Neunert, C. & Despotovic, J. Autoimmune hemolytic anemia and immune thrombocytopenia following hematopoietic stem cell transplant: a critical review of the literature. Pediatr. Blood Cancer 66, e27569 (2019).
Biasco, L. et al. In vivo tracking of human hematopoiesis reveals patterns of clonal dynamics during early and steady-state reconstitution phases. Cell Stem Cell 19, 107–119 (2016).
Busch, K. & Rodewald, H.-R. Unperturbed vs. post-transplantation hematopoiesis: both in vivo but different. Curr. Opin. Hematol. 23, 295 (2016).
Scala, S. et al. Dynamics of genetically engineered hematopoietic stem and progenitor cells after autologous transplantation in humans. Nat. Med. 24, 1683–1690 (2018).
Liu, P. et al. Bcl11a is essential for normal lymphoid development. Nat. Immunol. 4, 525–532 (2003).
Yu, Y. et al. Bcl11a is essential for lymphoid development and negatively regulates p53. J. Exp. Med. 209, 2467–2483 (2012).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168–e168 (2014).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Schafflick, D. et al. Integrated single cell analysis of blood and cerebrospinal fluid leukocytes in multiple sclerosis. Nat. Commun. 11, 247 (2020).
Acknowledgements
We thank all the patients and their families for their participation in the present study and the staff at Xiangya Hopital for their contributions to the care of the patients. This trial was sponsored by Shanghai Bioray Laboratories Inc. The sponsor was involved in clinical protocol design, data analysis, hematopoietic stem cell collection and cell production. We thank current and previous employees of Shanghai Bioray Laboratories Inc. and East China Normal University for helpful discussions. We thank S. Siwko for scientific editing and comments. We also thank D. E. Bauer for helpful suggestions to enhance our manuscript. We appreciate the support of grants from National Key R&D Program of China (grant nos. 2019YFA0802800, 2019YFA0110803, 2019YFA0109900 and 2019YFA0109901 to Y.W. and 2019YFA0802802 to M.L.), the Program for Professor of Special Appointment (Eastern Scholar) at Shanghai Institutions of Higher Learning (no. 11300-412214-20009 to Y.W.), the Innovation program of Shanghai Municipal Education Commission (no. 2019-01-07-00-05-E00054 to D.L.) and the Shanghai pujiang program (no. 11300-412213-19B08 to Y.W.).
Author information
Authors and Affiliations
Contributions
B.F., D.L., M.L. and Y.W. designed and led the study. B.F. was site principal investigator. J.L. and S.C. conducted experiments and analyzed data with the help of W.L., Q.W., F.Y., S.H., Y.J. and L.W. B.F., J.H. and F.C. were involved in patient care, testing and data presentation. B.F., J.L., S.C. and Y.W. wrote the manuscript. All authors contributed to the manuscript and approved its final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: W.L., F.Y., D.L., M.L. and Y.W. are employees of Bioray Laboratories. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Matthew Porteus, Martin Steinberg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling editor: Anna Maria Ranzoni, in collaboration with the Nature Medicine team
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Representative blood cell subtype cytometry analysis.
Representative blood cell subtype cytometry analysis. The gating strategy for blood cell subtype analysis is as shown. Lymphocytes were gated from a single cell population with CD45 high, SSC low. T cells are gated from the CD45+/CD3+ population. B cells are gated from the CD45+/CD3-/CD19+ population. NK cells are gated from the CD45+/CD3-/CD16/56+ population.
Extended Data Fig. 2 RP-HPLC analysis of globin chains in treated patients.
RP-HPLC analysis of globin chains in healthy donor and two patients (at Month 9, 15 and 18). RP-HPLC chromatograms are reported together with the non–α: α ratio (in brackets). The various globin chains detected were labeled on top and their respective peak shape ranges were marked with a gray background. Gamma globin chains were robustly induced in both patients. The expression of a common AγT chain variant was detected in samples derived from Patient 2. After treatment, the two patients had equal amounts of alpha and non-alpha chains in their blood after treatment as that of healthy donor.
Extended Data Fig. 3 Representative indel tables of two patients’ edited cells.
Summary of most frequent indels by deep sequencing of the genome from input CD34+ cells, PBMCs Mo7 after transplantation, and PBMCs Mo15 after transplantation from study patients. The reference sequence at the top of the figure is part of the BCL11A enhancer DNase I hypersensitive site (DHS) + 58 functional core. Gata1 motif marked with green box, and the sgRNA targets the complementary strand of the gata1 motif sequence.
Extended Data Fig. 4 Representative indel tables of two patients’ engrafted cells.
Summary of most frequent indels by deep sequencing result of T cells, B cells, NK cells, Monocytes and Neutrophils sorted from peripheral blood at Mo15 post transplantation from study patients. The reference sequence at the top of the figure is part of the BCL11A enhancer DNase I hypersensitive site (DHS) + 58 functional core. Gata1 motif marked with green box, and the sgRNA targets the complementary strand of the gata1 motif sequence.
Extended Data Fig. 5 Genotyping analysis of patient single CD34+ cell expansions.
CD34 + cells were sorted by MACS from bone marrow cells of patients 9 months after transplantation, and their single-cell expansions were analyzed for genotype by sanger sequencing. It was found that 88.3% (n=77) and 79.8% (n=104) of the CD34+ cells of the two patients were biallelic editing.
Extended Data Fig. 6 Representative cytoflow analysis for F-cells proportion in patient RBCs after treatment.
The proportion of F-cells in peripheral red blood cells was measured by FACS at 15 months after transplantation in both patients, blood sample from healthy people as control. After treatment, both patients showed robust HBF expression in peripheral red blood cells.
Extended Data Fig. 7 Sc-RNA seq analysis of edited and unedited patient PBMCs.
a, Marker gene expression for subtypes from PBMCs. b, UMAP plots representing 13 color-coded cell clusters identified in single-cell transcriptomes of PBMCs from 2 heathy donors, 1 patient sample before treatment and 3 patient samples at different time points after treatment. c, Violin plots showing BCL11A expression (log transformed) for B cell cluster from each sample. Wilcox two-tailed test, with each p-value, is indicated above the comparison group.
Supplementary information
Supplementary Information
Supplementary Tables 1–5, Clinical research protocol.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Rights and permissions
About this article
Cite this article
Fu, B., Liao, J., Chen, S. et al. CRISPR–Cas9-mediated gene editing of the BCL11A enhancer for pediatric β0/β0 transfusion-dependent β-thalassemia. Nat Med 28, 1573–1580 (2022). https://doi.org/10.1038/s41591-022-01906-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-01906-z
This article is cited by
-
Detection of CRISPR/Cas9-Mediated Fetal Hemoglobin Reactivation in Erythroblasts Derived from Cord Blood-Hematopoietic Stem Cells
Molecular Biotechnology (2024)
-
Recent advances in therapeutic CRISPR-Cas9 genome editing: mechanisms and applications
Molecular Biomedicine (2023)
-
Therapeutic adenine base editing of human hematopoietic stem cells
Nature Communications (2023)
-
Assessing and advancing the safety of CRISPR-Cas tools: from DNA to RNA editing
Nature Communications (2023)
-
CRISPR/Cas9, a promising approach for the treatment of β-thalassemia: a systematic review
Molecular Genetics and Genomics (2023)