Abstract
Richter syndrome (RS) arising from chronic lymphocytic leukemia (CLL) exemplifies an aggressive malignancy that develops from an indolent neoplasm. To decipher the genetics underlying this transformation, we computationally deconvoluted admixtures of CLL and RS cells from 52 patients with RS, evaluating paired CLL–RS whole-exome sequencing data. We discovered RS-specific somatic driver mutations (including IRF2BP2, SRSF1, B2M, DNMT3A and CCND3), recurrent copy-number alterations beyond del(9p21)(CDKN2A/B), whole-genome duplication and chromothripsis, which were confirmed in 45 independent RS cases and in an external set of RS whole genomes. Through unsupervised clustering, clonally related RS was largely distinct from diffuse large B cell lymphoma. We distinguished pathways that were dysregulated in RS versus CLL, and detected clonal evolution of transformation at single-cell resolution, identifying intermediate cell states. Our study defines distinct molecular subtypes of RS and highlights cell-free DNA analysis as a potential tool for early diagnosis and monitoring.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
WES, RNA-seq, WGS, scRNA-seq and cfDNA data are available at dbgap (https://www.ncbi.nlm.nih.gov/gap/) using accession number phs002458.v2.p1 (http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs002458.v2.p1). RNA-seq data from validation cohort can be accessed by registering for an EGA account (https://ega-archive.org/) and contacting the Data Access Committee under study EGAS00001005495 and accession number EGAD00001007922. The following figures have associated raw data: Fig. 2b–h, Fig. 4a,f, Fig. 5a,b,d,e, Fig. 6g, Extended Data Fig. 4a–x, Extended Data Fig. 5a–c, Extended Data Fig. 6a, Extended Data Fig. 8, Extended Data Fig. 9, and Extended Data Fig. 10.
Code availability
Code is available for CNVSingle (https://github.com/broadinstitute/CNVsingle) and PhylogicNDT CopyNumber2Tree (https://github.com/broadinstitute/PhylogicNDT).
References
Offin, M. et al. Concurrent RB1 and TP53 alterations define a subset of EGFR-mutant lung cancers at risk for histologic transformation and inferior clinical outcomes. J. Thorac. Oncol. 14, 1784–1793 (2019).
Volta, A. D. et al. Transformation of prostate adenocarcinoma into small-cell neuroendocrine cancer under androgen deprivation therapy: much is achieved but more information is needed. J. Clin. Oncol. 37, 350–351 (2019).
Parikh, S. A., Kay, N. E. & Shanafelt, T. D. How we treat Richter syndrome. Blood 123, 1647–1657 (2014).
Landau, D. A. et al. Mutations driving CLL and their evolution in progression and relapse. Nature 526, 525–530 (2015).
Puente, X. S. et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature 526, 519–524 (2015).
Knisbacher, B. A. et al. Molecular map of chronic lymphocytic leukemia and its impact on outcome. Nat. Genet. 54, 1664–1674 (2022).
Chigrinova, E. et al. Two main genetic pathways lead to the transformation of chronic lymphocytic leukemia to Richter syndrome. Blood 122, 2673–2682 (2013).
Fabbri, G. et al. Genetic lesions associated with chronic lymphocytic leukemia transformation to Richter syndrome. J. Exp. Med. 210, 2273–2288 (2013).
Klintman, J. et al. Genomic and transcriptomic correlates of Richter transformation in chronic lymphocytic leukemia. Blood 137, 2800–2816 (2021).
Rossi, D. et al. The genetics of Richter syndrome reveals disease heterogeneity and predicts survival after transformation. Blood 117, 3391–3401 (2011).
Taylor-Weiner, A. et al. DeTiN: overcoming tumor-in-normal contamination. Nat. Methods 15, 531–534 (2018).
Leshchiner, I. et al. Comprehensive analysis of tumour initiation, spatial and temporal progression under multiple lines of treatment. Preprint at bioRxiv 508127 (2018).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Schmitz, R. et al. Genetics and pathogenesis of diffuse large B-cell lymphoma. N. Engl. J. Med. 378, 1396–1407 (2018).
Chapuy, B. et al. Genomic analyses of PMBL reveal new drivers and mechanisms of sensitivity to PD-1 blockade. Blood 134, 2369–2382 (2019).
Biran, A. et al. Activation of Notch and Myc Signaling via B-cell-restricted depletion of Dnmt3a generates a consistent murine model of chronic lymphocytic leukemia. Cancer Res. 81, 6117–6130 (2021).
Mahajan, V. S. et al. B1a and B2 cells are characterized by distinct CpG modification states at DNMT3A-maintained enhancers. Nat. Commun. 12, 2208 (2021).
Chapuy, B. et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat. Med. 24, 679–690 (2018).
Challa-Malladi, M. et al. Combined genetic inactivation of beta2-Microglobulin and CD58 reveals frequent escape from immune recognition in diffuse large B cell lymphoma. Cancer Cell 20, 728–740 (2011).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
Gettinger, S. et al. Impaired HLA Class I antigen processing and presentation as a mechanism of acquired resistance to immune checkpoint inhibitors in lung cancer. Cancer Disco. 7, 1420–1435 (2017).
Singh, K. et al. c-MYC regulates mRNA translation efficiency and start-site selection in lymphoma. J. Exp. Med. 216, 1509–1524 (2019).
Lee, S. C. et al. Synthetic lethal and convergent biological effects of cancer-associated spliceosomal gene mutations. Cancer Cell 34, 225–241 e228 (2018).
Edelmann, J. et al. Genomic alterations in high-risk chronic lymphocytic leukemia frequently affect cell cycle key regulators and NOTCH1-regulated transcription. Haematologica 105, 1379–1390 (2020).
Anderson, M. A. et al. Clinicopathological features and outcomes of progression of CLL on the BCL2 inhibitor venetoclax. Blood 129, 3362–3370 (2017).
Jain, P. et al. Long-term outcomes for patients with chronic lymphocytic leukemia who discontinue ibrutinib. Cancer 123, 2268–2273 (2017).
Maddocks, K. J. et al. Etiology of ibrutinib therapy discontinuation and outcomes in patients with chronic lymphocytic leukemia. JAMA Oncol. 1, 80–87 (2015).
Burger, J. A. et al. Clonal evolution in patients with chronic lymphocytic leukaemia developing resistance to BTK inhibition. Nat. Commun. 7, 11589 (2016).
Guieze, R. et al. Mitochondrial reprogramming underlies resistance to BCL-2 inhibition in lymphoid malignancies. Cancer Cell 36, 369–384 e313 (2019).
Kasar, S. et al. Whole-genome sequencing reveals activation-induced cytidine deaminase signatures during indolent chronic lymphocytic leukaemia evolution. Nat. Commun. 6, 8866 (2015).
Gruber, M. et al. Growth dynamics in naturally progressing chronic lymphocytic leukaemia. Nature 570, 474–479 (2019).
Zhang, N. et al. Overexpression of Separase induces aneuploidy and mammary tumorigenesis. Proc. Natl Acad. Sci. USA 105, 13033–13038 (2008).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Adalsteinsson, V. A. et al. Scalable whole-exome sequencing of cell-free DNA reveals high concordance with metastatic tumors. Nat. Commun. 8, 1324 (2017).
Nadeu, F. et al. Detection of early seeding of Richter transformation in chronic lymphocytic leukemia. Nat. Med. 28, 1662–1671 (2022).
Miller, C. R. et al. Near-tetraploidy is associated with Richter transformation in chronic lymphocytic leukemia patients receiving ibrutinib. Blood Adv. 1, 1584–1588 (2017).
Quinton, R. J. et al. Whole-genome doubling confers unique genetic vulnerabilities on tumour cells. Nature 590, 492–497 (2021).
Soilleux, E. J. et al. Diagnostic dilemmas of high-grade transformation (Richter’s syndrome) of chronic lymphocytic leukaemia: results of the phase II National Cancer Research Institute CHOP-OR clinical trial specialist haemato-pathology central review. Histopathology 69, 1066–1076 (2016).
Favini, C. et al. Clonally unrelated Richter syndrome are truly de novo diffuse large B-cell lymphomas with a mutational profile reminiscent of clonally related Richter syndrome. Br. J. Haematol. 198, 1016–1022 (2022).
Parikh, A. R. et al. Liquid versus tissue biopsy for detecting acquired resistance and tumor heterogeneity in gastrointestinal cancers. Nat. Med. 25, 1415–1421 (2019).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303 (2010).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Gao, G. F. et al. Tangent normalization for somatic copy-number inference in cancer genome analysis. Bioinformatics 38, 4677–4686 (2022).
Consortium, I.T.P.-C.A.o.W.G. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222 (2016).
Wala, J. A. et al. SvABA: genome-wide detection of structural variants and indels by local assembly. Genome Res 28, 581–591 (2018).
Bass, A. J. et al. Genomic sequencing of colorectal adenocarcinomas identifies a recurrent VTI1A-TCF7L2 fusion. Nat. Genet. 43, 964–968 (2011).
Drier, Y. et al. Somatic rearrangements across cancer reveal classes of samples with distinct patterns of DNA breakage and rearrangement-induced hypermutability. Genome Res 23, 228–235 (2013).
Sondka, Z. et al. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nat. Rev. Cancer 18, 696–705 (2018).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Brunet, J. P., Tamayo, P., Golub, T. R. & Mesirov, J. P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl Acad. Sci. USA 101, 4164–4169 (2004).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Zhang, Y., Parmigiani, G. & Johnson, W. E. ComBat-seq: batch effect adjustment for RNA-seq count data. NAR Genom. Bioinform 2, lqaa078 (2020).
Monti, S., Tamayo, P., Mesirov, J. & Golub, T. Consensus clustering: a resampling-based method for class discovery and visualization of gene expression microarray data. Mach. Learn. 52, 91–118 (2003).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 e324 (2019).
Acknowledgements
We thank C. Hahn, E. Ten Hacken, W. Zhang, S. Gohil and L. Werner for helpful discussions. We thank C. Patterson, S. Pollock, O. Olive, C. J. Shaughnessy, F. Dao and H. Lyon for assistance in data collection and organization and S. Belkin and C. Birger for assistance in data storage. We thank T. Lehmberg, M. McDonough, C. Galler and M. Collins for assistance in sample collection and biobanking. We thank the patients, their families and the investigators of the clinical trials for providing samples and clinical data. This study was supported by NIH/NCI P01 CA206978 (to C.J.W. and G.G.) and NCI (1U10CA180861-01) (to C.J.W.). The work is partially supported by the Broad/IBM Cancer Resistance Research Project (I.L., G.G. and L.P.) and a grant from Force Hemato (R.G.). Individual support was provided by DDCF Physician-Scientist Fellowship (E.M.P.), Dana-Farber Flames FLAIR fellowship (E.M.P.), ASCO Conquer Cancer Young Investigator Award (E.M.P.), Fishman Family Fund (R.G. and C.L.), EMBO fellowship ALTF 14-2018 (B.A.K.), NCI Research Specialist Award R50CA251956 (S.L.) and NIH/NCI R21CA267527-01 (S.Y.). Additional research support was provided by NIH R01 CA213442 (J.R.B.), Melton Family Foundation (J.R.B.), NIH/NCI R01-CA236361 (T.J.K.) and the Deutsche Forschungsgemeinschaft (DFG) SFB1074 subprojects B10 (E.T.) and subprojects B1 and B2 (E.T., C.S. and S.S.)
Author information
Authors and Affiliations
Contributions
E.M.P., I.L., R.G., C.J., G.G., S.S. and C.J.W. designed and performed the experiments, analyzed the data and wrote the manuscript. E.M.P., N.P.-Z., A.J.A., T.H. and S.L. generated single-cell RNA-seq data. C. Lemvigh, E.M.P. and I.L. analyzed single-cell RNA-seq data, along with C. Levovitz, F.U. and K.R. R.G., E.T., C. Schneider, M.S.D., N.J., W.W., L.R., T.J.K, J.B., S.H., P.F., F.C., N.K., S.P., J.R.B. and S.S. provided patient samples. K.J.L. and S.L. designed targeted NGS assay for detecting the NOTCH1 3′-UTR mutation. N.R. performed mapping and analysis of these NGS data. D.R., F.U., C. Levovitz and S.Y. analyzed the RNA-seq data. S.D. analyzed the mutational data under the supervision of E.M.P. and C.J.W. C.M., J.M., J.H., L.L. and C. Stewart analyzed the WGS data. C.J., I.L., B.P., L.E., D.R., A.T-W., A.M., D.L., E.M.P., R.G. and C.J.W. performed sequencing data analyses, assessment of the clonal architecture and inference of phylogenies under the supervision of I.L. and G.G. E.M.P., R.G., L.R., J.B. and S.F. prepared patient samples. I.L. developed the analytic tool for determining somatic copy-number variations from FFPE samples and CNVSingle for detecting copy-number events in single-cell RNA-seq data. D.N. performed and supervised statistical analyses. R.R. performed statistical analyses. G.F. provided graphical representation of the clinical data. B.A.K. performed immunogenetic data analyses. B.P.D., K.R., C. Levovitz and L.P. helped to design and guide the research. B.P.D., R.A.J. and K.S. were involved in managing the project. I.L., C.J. and Z.L. performed cell-free DNA analyses. All authors discussed, interpreted results and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
I.L. serves as a consultant for PACT Pharma, Inc.; has stock in, is on the board of and serves as a consultant for ennov1 LLC.; and is on the board of and holds equity in Nord Bio, Inc. C.J.W., G.G., B.A.K. and Z.L. are inventors on a patent: ‘Compositions, Panels, and Methods for Characterizing Chronic Lymphocytic Leukemia’ (PCT/US21/45144). C.J.W., G.G., E.M.P., I.L. and R.G. are named as inventors on U.S. provisional patent application serial number 63/244,625, filed on 15 September 2021, and U.S. provisional patent application serial number 63/291,213, filed on 17 December 2021, both of which are entitled ‘Diagnosis and Prognosis of Richter’s Syndrome.’ G.G. is a founder of and consultant for and holds privately held equity in Scorpion Therapeutics; receives funding support from IBM and Pharmacyclics; and is an inventor on patent applications related to MSMuTect, MSMutSig, MSIDetect, POLYSOLVER and SignatureAnalyzer-GPU. R.G. receives funding support from Abbvie, AstraZeneca, Gilead, Janssen and Roche. M.S.D. served as a consultant for Abbvie, Adaptive Biotechnologies, Ascentage Pharma, AstraZeneca, BeiGene, Bristol-Myers Squibb, Eli Lilly, Genentech/Roche, Janssen, Merck, Ono Pharmaceuticals, Pharmacyclics, Research to Practice, Takeda, TG Therapeutics, Verastem and Zentalis; receives funding support from Ascentage Pharma, AstraZeneca, Genentech/Roche, MEI Pharma, Novartis, Pharmacyclics, Surface Oncology, TG Therapeutics and Verastem; and receives funding for travel from Abbvie, BeiGene, BioAscend, Clinical Care Options, Curio Science, Imedex, ION Solutions, Janssen, MDOutlook, PeerView, PRIME Oncology, Research to Practice and TG Therapeutics. J.R.B. has served as a consultant for Abbvie, Acerta/AstraZeneca, BeiGene, Bristol-Myers Squibb/Juno/Celgene, Catapult, Eli Lilly, Genentech/Roche, Hutchmed, Janssen, MEI Pharma, MorphoSys AG, Novartis, Pfizer, Pharmacyclics and Rigel; and received research funding from Gilead, Loxo/Lilly, SecuraBio, Sun and TG Therapeutics. C.J.W. receives funding support from Pharmacyclics and holds equity in BioNTech, Inc. N.E.K. serves as an advisor for Abbvie, AstraZeneca, Beigene, Behring, Cytomx Therapy, Dava Oncology, Janssen, Juno Therapeutics, Oncotracker, Pharmacyclics and Targeted Oncology; receives funding support from Abbvie, Acerta Pharma, Bristol Meyer Squib, Celgene, Genentech, MEI Pharma, Pharmacyclics, Sunesis, TG Therapeutics and Tolero Pharmaceuticals; and participates on the Data Safety Monitoring Committee for Agios Pharm, AstraZeneca, BMS-Celgene, Cytomx Therapeutics, Dren Bio, Janssen, MorphoSys and Rigel. T.J.K. is on the advisory board and receives funding support from Abbvie and Roche and serves on the speakers’ bureau for Abbvie, Janssen and Roche. E.T. serves as an advisor and is on the speakers’ bureau for Abbvie, Janssen and Roche; and receives funding support from Abbvie and Roche. S.S. is on the advisory board and receives funding, travel support and speakers’ fees from AbbVie, AstraZeneca, BeiGene, BMS, Celgene, Gilead, GSK, Hoffmann-La Roche, Janssen, Novartis and Sunesis. N.J. receives research funding from AbbVie, Adaptive Biotechnologies, ADC Therapeutics, Aprea Therapeutics, AstraZeneca, BMS, Cellectis, Fate Therapeutics, Genentech, Incyte, Loxo Oncology, Medisix, Mingsight, Novalgen, Pfizer, Pharmacyclics, Precision BioSciences, Servier and Takeda; and serves on the advisory board/receives honoraria from AbbVie, Adaptive Biotechnologies, ADC Therapeutics, AstraZeneca, Beigene, BMS, Cellectis, Genentech, Janssen, MEI Pharma, Pharmacyclics, Precision BioSciences, Servier and TG Therapeutics. W.G.W. reports funding from Abbvie, AstraZeneca/Acerta Pharma, Cyclacel, Genentech, Gilead Sciences, GSK/Novartis, Janssen, Juno Therapeutics, KITE Pharma, Loxo Oncology, Inc., Miragen, Oncternal Therapeutics, Inc., Pharmacyclics LLC, Sunesis and Xencor. S.A.P. has received research funding to the institution from AbbVie, Ascentage Pharma, AstraZeneca, Janssen, Merck, Pharmacyclics and TG Therapeutics for clinical studies in which S.A.P. is a principal investigator. S.A.P has received honoraria for participation in consulting activities/advisory board meetings for AbbVie, Adaptive Biotechnologies, Amgen, AstraZeneca, Genentech, GlaxoSmithKline, Merck and Pharmacyclics (no personal compensation). K.J.L. holds equity in Standard BioTools, Inc. D.N. has stock ownership in Madrigal Pharmaceuticals. C.S. serves on a speakers’ bureau for AbbVie and AstraZeneca. D.L. holds stock in and consults for ennov1. N.P is currently an employee at Bristol Meyers Squibb. All remaining authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Daniel Hodson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Anna Maria Ranzoni, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
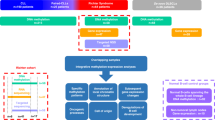
Extended Data Fig. 1 Clonal deconvolution process.
a, distinguishing RS from CLL clones after inferring subclonal composition of paired CLL and RS samples. b, inferring phylogenetic tree from cancer cell fraction using PhylogicNDT. c, sample composition d, mapping copy-number variations to clones using CopyNumber2Tree.
Extended Data Fig. 2 Phylogenetic reconstruction and somatic genomic alterations.
For each of the patient trios with WES data, the left panel shows the phylogenetic tree tracing the transformation history from CLL to RS. The magenta frame denotes the Richter clones. The middle top panel represents the subclonal composition inferred after clustering alterations with similar cancer cell fractions as previously reported4. The middle bottom panel indicates the timeline with RS and CLL sampling time and CLL therapeutic lines. (F, fludarabine; C, cyclophosphamide; R, rituximab; P, pentostatin; O/Ofa, ofatumumab; HDMP, high-dose methylprednisolone; A, alemtuzumab; Auto, autologous stem cell transplantation; CLB, chlorambucil; B, bendamustine; CHOP, cyclophosphamide, doxorubicin, vincristine, prednisone; ESHAP, etoposide, methylprednisolone, high-dose cytarabine, cisplatin; CHP, cyclophosphamide, doxorubicin, prednisone; Len, lenalidomide; Ob, obinutuzumab; idela; idelalisisb; D, dexamethasone; Adria, adriamycin). The right panel is composed of allelic fraction plots and allelic copy ratio plots showing clonal assignment of somatic copy-number events to CLL and RS clones. Cases with whole genome doubling in Extended Data Fig. 2 and clonal unrelated cases in Extended Data Fig. 3.
Extended Data Fig. 3 Phylogenetic reconstruction and somatic genomic alterations.
For each of the patient trios with WES data, the left panel shows the phylogenetic tree tracing the transformation history from CLL to RS. The magenta frame denotes the Richter clones. The middle top panel represents the subclonal composition inferred after clustering alterations with similar cancer cell fractions as previously reported4. The middle bottom panel indicates the timeline with RS and CLL sampling time and CLL therapeutic lines. (F, fludarabine; C, cyclophosphamide; R, rituximab; P, pentostatin; O/Ofa, ofatumumab; HDMP, high-dose methylprednisolone; A, alemtuzumab; Auto, autologous stem cell transplantation; CLB, chlorambucil; B, bendamustine; CHOP, cyclophosphamide, doxorubicin, vincristine, prednisone; ESHAP, etoposide, methylprednisolone, high-dose cytarabine, cisplatin; CHP, cyclophosphamide, doxorubicin, prednisone; Len, lenalidomide; Ob, obinutuzumab; idela; idelalisisb; D, dexamethasone; Adria, adriamycin). The right panel is composed of allelic fraction plots and allelic copy ratio plots showing clonal assignment of somatic copy-number events to CLL and RS clones. Cases with whole genome doubling in Extended Data Fig. 2 and clonal unrelated cases in Extended Data Fig. 3.
Extended Data Fig. 4 Putative RS driver genes.
a-x, individual protein mutation maps for selected putative Richter drivers, showing gene mutation subtype (for example, missense), position and evidence of mutational hotspots. Panels were generated by using the cBioPortal for Cancer Genomics tool.
Extended Data Fig. 5 RS sCNAs and genomic clustering.
GISTIC2-defined recurrent copy-number gains (red, left) and losses (blue, right) are visualized for focal events for RS samples (a) and RS clones (b) (RS samples with CLL events subtracted, bottom). Chromosomes are shown on the vertical axis. Green line denotes a near significant q value of 0.25 and significant events (q<0.1) are annotated in text along with putative driver genes contained within the peak (Supplementary Table 5) c, NMF clustering of RS with DLBCL (304 de novo DLBCL samples19 shows clonal related RS clusters separately from DLBCL and closes to DLBCL from C2 (ref. 19). Clonal unrelated RS clusters across DLBCL subtypes and separate from RS. Samples were annotated for clonal relationship (related RS, gray, unrelated RS, black), cohort (DLBCL, light purple; RS, dark purple) and DLBCL clusters (C1, purple; C2, yellow, C3, pink, C4, blue, C5, green)19. d, NMF clustering of RS shows 5 distinct genomic subtypes of transformation.
Extended Data Fig. 6 Transcriptome supports distinct RS molecular subtypes.
a, Supervised clustering of transcriptome data from 36 RS patients by molecular subtype highlights differentially regulated genes in subtype 1 and 3 (Supplementary Table 8). Samples are annotated for cohort (Discovery, pink; Validation, yellow), clonal relationship (unrelated, black, related, white), and sample purity by WES (green gradient). b, Unsupervised consensus clustering of RS transcriptome data (n=36) shows 5 clusters. (Discovery, pink; Validation, yellow), RS molecular subtype (1, purple; 2, blue; 3, orange; 4, green; and 5, pink), and sample purity by WES (green gradient).c, 5 × 5 table showing association between molecular subtype of RS and unsupervised transcriptome clusters (2 sided Fisher’s exact test, P=0.038) d, Kaplan–Meier curve showing OS of clonal unrelated RS compared to clonal related RS. P value is log rank (2 sided Mantel Cox).
Extended Data Fig. 7 Phylogenetic trees showing CLL and RS clones from WGS of paired samples.
a, Phylogenetic tree and CCF plot for 9 patients based on WGS data showing clonal related RS (magenta box). b, Phylogenetic tree and CCF plot for 2 patients based on WGS demonstrating clonally unrelated RS c, Representative phylogenetic trees and CCF plot for 3 patients from UK cohort9 based on WGS.
Extended Data Fig. 8 WGS Circos plots with or without chromothripsis.
a, chromothripsis and kataegis in RS sample (Pt 42) with whole genome doubling. Circos plots showing structural variants (interchromosomal, blue; deletion, red; inversion, yellow; tandem duplication, green; long range, teal), allelic copy number (middle), rainfall plot with kataegis regions (red) and chromosomes (outside). Adjacent rainfall plots show kategis regions (C to G, red; C to T, yellow; C to A, teal) with corresponding allelic copy-number fragmentation. b, Circos plots from RS WGS samples showing structural variants (interchromosomal, blue; deletion, red; inversion, yellow; tandem duplication, green and long range, teal), allelic copy number (middle), rainfall plot with kataegis regions (red) and chromosomes (outside). SVs impacting known genes and translocation partners are labeled (Supplementary Table 7k).
Extended Data Fig. 9 Single cell processing and transcriptome analysis of RS samples at single cell resolution.
a, flow sorting strategy for RS single-cell samples. Flow sorting to separate RS and CLL cells by size for Patient 19 and Patient 41 (lymph node, LN; peripheral blood, PB; bone marrow, BM). Flow sorting viable cells for Pt 43, Pt 4 and Pt10. Representative flow plots below demonstrate CLL and RS cells were included in sorted population. b, B cell receptor (BCR) clonotypes plotted for RS and CLL clusters on UMAP visualization. c, Representative example from patient 10 showing CNVsingle identifies malignant B cell clusters (5 and 6) separate from immune cell clusters (0,1,2,3,4,7,9). d, UMI/cell and Gene/cell plots for CLL and RS single-cell clusters. RS demonstrates higher UMI/cell (P<2.2 × 10–16 see Methods, Supplementary Table 8). e, RNA inference of directional trajectories is shown on UMAP visualization for Pts 43 and 10. f, copy-number variation heatmap inferred in each cluster from scRNA-seq data using our CNVSingle algorithm for Pts 43 and 10 (Methods).
Extended Data Fig. 10 Single-cell transcriptome and copy-number analysis of RS patients.
a, UMAP visualization of single-cells from patient 4 (left) with associated allelic copy-number ratio plot inferred by CNVsingle (top right) and RS WES (bottom right). b, UMAP visualization of CLL and RS cells from Patient 18 (left top panel) with flow-sorting annotations (right top panel). Inferred CNAs from CNVSingle (bottom panel) are shown as heatmap with CLL (green) and RS (pink) events highlighted. c, UMAP visualization of CLL and RS cells from Patient 41 (left top panel) with flow-sorting annotations (right top panel). Inferred CNAs from CNVSingle (bottom panel) with CLL (green) and RS (pink) events highlighted. d, Plasma of patient 44 shows RS-specific sCNVs on chromosome 9 and 13 leading up to RS diagnosis, which are not reflected in circulating CLL e, Plasma of patient 99 at the start of CLL-directed therapy (top) and just ahead of diagnosis of RS (bottom) during CLL response. f, Chromothripsis in post-transplant RS plasma cfDNA at time of relapse (Pt 112). g, Plot showing allele frequency of RS (purple) and CLL (green) mutations in RS WES (bottom) and plasma cfDNA WES (top) for patient 5 (top) and patient 44 (bottom).
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Tables
Supplementary Tables 1–10 combined into single Excel document.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parry, E.M., Leshchiner, I., Guièze, R. et al. Evolutionary history of transformation from chronic lymphocytic leukemia to Richter syndrome. Nat Med 29, 158–169 (2023). https://doi.org/10.1038/s41591-022-02113-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-02113-6
This article is cited by
-
NFKBIE mutations are selected by the tumor microenvironment and contribute to immune escape in chronic lymphocytic leukemia
Leukemia (2024)
-
Chronic lymphocytic leukemia patient-derived xenografts recapitulate clonal evolution to Richter transformation
Leukemia (2024)
-
Treatment of Richter’s Transformation with Novel Therapies
Current Hematologic Malignancy Reports (2024)
-
Allogeneic hematopoietic stem-cell transplantation for patients with Richter transformation: a retrospective study on behalf of the Chronic Malignancies Working Party of the EBMT
Bone Marrow Transplantation (2024)
-
Tislelizumab plus zanubrutinib for Richter transformation: the phase 2 RT1 trial
Nature Medicine (2024)